This examine describes the characterization of the lately described Salmonella genomic island 1 (SGI1) (D. A. Boyd, G. A. Peters, L.-Okay. Ng, and M. R. Mulvey, FEMS Microbiol. Lett.
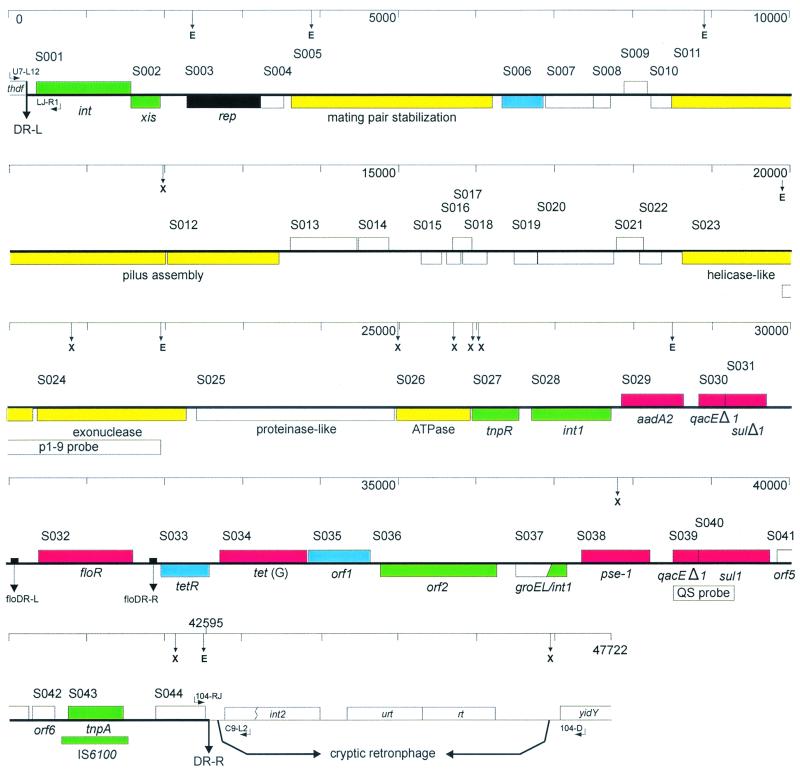
189:285-291, 2000), which harbors the genes associated with the ACSSuT phenotype in a Canadian isolate of Salmonella enterica serovar Typhimurium DT104. A 43-kb region has been utterly sequenced and discovered to comprise 44 predicted open studying frames (ORFs) which comprised roughly 87% of the whole sequence. Fifteen ORFs didn’t present any vital homology to identified gene sequences.
A quantity of ORFs present vital homology to plasmid-related genes, suggesting, a minimum of in half, a plasmid origin for the SGI1, though some with homology to phage-related genes had been recognized.
The SGI1 was recognized in a quantity of multidrug-resistant DT120 and S. enterica serovar Agona strains with comparable antibiotic-resistant phenotypes. The G+C content material suggests a potential mosaic construction for the SGI1. Emergence of the SGI1 in serovar Agona strains is mentioned.
A 300 kilobase interval genetic map of rice together with 883 expressed sequences.
We have constructed a excessive decision rice genetic map containing 1,383 DNA markers at a median interval of 300 kilobases (kb). The markers, distributed alongside 1,575 cM on 12 linkage teams, comprise 883 cDNAs, 265 genomic DNAs, 147 randomly amplified polymorphic DNAs (RAPD) and 88 different DNAs.
cDNAs had been derived from rice root and callus, analysed by single-run sequencing and looked for similarities with identified proteins. Nearly 260 rice genes are newly recognized and mapped, and genomic DNA and cloned RAPD fragments had been additionally sequenced to generate STSs.
Our map is the first vital gene expression map in crops. It can be the densest genetic map out there in crops and the first to be backed up comprehensively by clone sequence information.
